 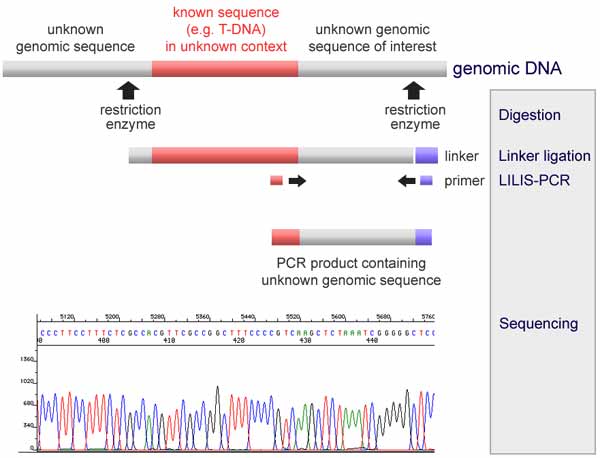
MacVectorTip: Identifying, Selecting and Assembling NGS reads with a variant genotype.Run FILE | NEW | ASSEMBLY PROJECT, Choose a reference sequence, add your sequencing reads and also any reference sequences. Choose a reference sequence, then add your trace files from the sequencing facility and click ALIGN. Run FILE | NEW | ALIGN SEQUENCES TO A REFERENCE. To align a small sequencing project against a reference. Right click (or CTRL+left Click) on any faded ORF or Feature on your sequence. Double click the hit to open it directly in MacVector. 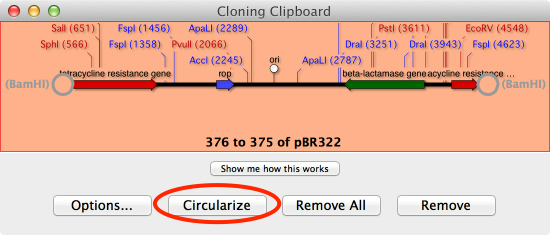
Select DATABASE | ONLINE KEYWORD SEARCH (ENTREZ), enter the accession number of your favorite gene and hit search. Open two genomes and run ANALYZE | COMPARE GENOMES BY FEATURE. List genetic differences between one organism and another related strain Select FILE | NEW | ALIGNMENT, Click ADD SEQS and then ALIGN. Select FILE | NEW | AGAROSE GEL, Drag a restriction site from the Map tab of a sequence and drop on the Agarose Gel window Type or paste in your forward primer, Repeat for the reverse primer. For the left primer click USE THIS Primer. Open your template sequence, Go to ANALYZE | PRIMER DESIGN/TEST(PAIRS). Add a restriction site or mutation, slide the primer until the oligo is optimal. Gibson Assembly/Ligase Independent CloningįILE | NEW | GIBSON/LIGASE INDEPENDENT ASSEMBLY. Choose the type of project, add a cloning vector, then drag a gene from a sequence to clone. Drag the digested fragment from the Cloning Clipboard to your vector and click LIGATE. Select two restriction sites and click DIGEST. Here’s an overview of workflows in MacVector for the molecular biologist. MacVector has a wide array of different tools for working with protein and DNA sequences. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
